MicroRNAs (miRNAs) are post-transcriptional regulators that bind to complementary sequences in the three prime untranslated regions (3' UTRs) of target messenger RNA transcripts (mRNAs), usually resulting in gene silencing.[1][2] miRNAs are short ribonucleic acid (RNA) molecules, on average only 22 nucleotides long. The human genome may encode over 1000 miRNAs,[3][4] which may target about 60% of mammalian genes[5] and are abundant in many human cell types.[6] Each miRNA may repress hundreds of mRNAs.[7][8] MiRNAs are well conserved in eukaryotic organisms and are thought to be a vital and evolutionarily ancient component of genetic regulation.[9][10][11][12][13]
The first miRNAs were characterized in the early 1990s, but miRNAs were not recognized as a distinct class of biologic regulators with conserved functions until the early 2000s. Since then, miRNA research has revealed multiple roles in negative regulation (transcript degradation and sequestering, translational suppression) and possible involvement in positive regulation (transcriptional and translational activation). By affecting gene regulation, miRNAs are likely to be involved in most biologic processes.[14][15][16][17][18][19][20][21] Different sets of expressed miRNAs are found in different cell types and tissues.[22]
Aberrant expression of miRNAs has been implicated in numerous disease states, and miRNA-based therapies are under investigation.[23][24][25]
History
MicroRNAs were discovered in 1993 by Victor Ambros, Rosalind Lee and Rhonda Feinbaum during a study of the gene lin-14 in C. elegans development.[26] They found that LIN-14 protein abundance was regulated by a short RNA product encoded by the lin-4 gene. A 61 nucleotide precursor from lin-4 gene matured to a 22 nucleotide RNA containing sequences partially complementary to multiple sequences in the 3’ UTR of the lin-14 mRNA. This complementarity was sufficient and necessary to inhibit the translation of lin-14 mRNA into LIN-14 protein. Retrospectively, the lin-4 small RNA was the first microRNA to be identified, though at the time, it was thought to be a nematode idiosyncrasy. Only in 2000 was a second RNA characterized: let-7, which repressed lin-41, lin-14, lin-28, lin-42, and daf-12 expression during developmental stage transitions in C. elegans. let-7 was soon found to be conserved in many species,[27][28] indicating the existence of a wider phenomenon.
Nomenclature
Under a standard nomenclature system, names are assigned to experimentally confirmed miRNAs before publication of their discovery.[29][30] The prefix "mir" is followed by a dash and a number, the latter often indicating order of naming. For example, mir-123 was named and likely discovered prior to mir-456. The uncapitalized "mir-" refers to the pre-miRNA, while a capitalized "miR-" refers to the mature form. miRNAs with nearly identical sequences bar one or two nucleotides are annotated with an additional lower case letter. For example, miR-123a would be closely related to miR-123b. miRNAs that are 100% identical but are encoded at different places in the genome are indicated with additional dash-number suffix: miR-123-1 and miR-123-2 are identical but are produced from different pre-miRNAs. Species of origin is designated with a three-letter prefix, e.g., hsa-miR-123 would be from human (Homo sapiens) and oar-miR-123 would be a sheep (Ovis aries) miRNA. Other common prefixes include 'v' for viral (miRNA encoded by a viral genome) and 'd' for Drosophila miRNA (a fruit fly commonly studied in genetic research). microRNAs originating from the 3’ or 5’ end of a pre-miRNA are denoted with a -3p or -5p suffix. (In the past, this distinction was also made with 's' (sense) and 'as' (antisense)). When relative expression levels are known, an asterisk following the name indicates an miRNA expressed at low levels relative to the miRNA in the opposite arm of a hairpin. For example, miR-123 and miR-123* would share a pre-miRNA hairpin, but relatively more miR-123 would be found in the cell.
Biogenesis
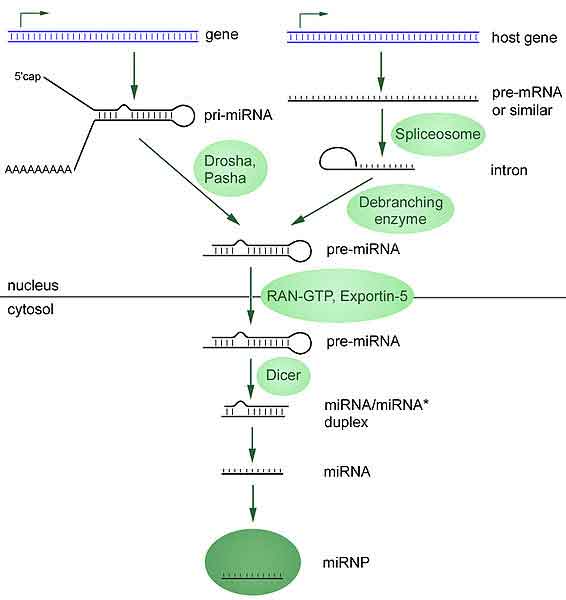
MicroRNAs are produced from either their own genes or from introns (*)
Most microRNA genes are found in intergenic regions or in anti-sense orientation to genes[31] and contain their own miRNA gene promoter and regulatory units.[31][32][33][34] As much as 40% of miRNA genes may lie in the introns of protein and non-protein coding genes or even in exons.[35] These are usually, though not exclusively, found in a sense orientation [36][37] and thus usually are regulated together with their host genes.[38][39][40] Other miRNA genes showing a common promoter include the 42-48% of all miRNAs originating from polycistronic units contaning 2-7 discrete loops from which mature miRNAs are processed,[41][32] although this does not necessarily mean the mature miRNAs of a family will be homologous in structure and function. The promoters mentioned have been shown to have some similarities in their motifs to promoters of other genes transcribed by RNA polymerase II such as protein coding genes.[32][42] The DNA template is not the final word on mature miRNA production: 6% of human miRNAs show RNA editing, the site-specific modification of RNA sequences to yield products different from those encoded by their DNA. This increases the diversity and scope of miRNA action beyond that implicated from the genome alone.
Transcription
miRNA genes are usually transcribed by RNA polymerase II (Pol II).[32][43] The polymerase often binds to a promoter found near the DNA sequence encoding what will become the hairpin loop of the pre-miRNA. The resulting transcript is capped with a specially-modified nucleotide at the 5’ end, polyadenylated with multiple adenosines (a poly(A) tail),[36][32] and spliced. The product, called a primary miRNA (pri-miRNA), may be hundreds or thousands of nucleotides in length and contain one or more miRNA stem loops.[36][32] When a stem loop precursor is found in the 3’ UTR, a transcript may serve as a pri-miRNA and a mRNA.[36] RNA polymerase III (Pol III) transcribes some miRNAs, especially those with upstream Alu sequences, transfer RNAs (tRNAs), and mammalian wide interspersed repeat (MWIR) promoter units.[44]
Nuclear processing
A single pri-miRNA may contain from one to six miRNA precursors. These hairpin loop structures are composed of about 70 nucleotides each. Each hairpin is flanked by sequences necessary for efficient processing. The double-stranded RNA structure of the hairpins in a pri-miRNA is recognized by a nuclear protein known as DiGeorge Syndrome Critical Region 8 (DGCR8 or "Pasha" in invertebrates), named for its association with DiGeorge Syndrome. DGCR8 associates with the enzyme Drosha, a protein that cuts RNA, to form the "Microprocessor" complex.[45] In this complex, DGCR8 orients the catalytic RNase III domain of Drosha to liberate hairpins from pri-miRNAs by cleaving RNA about eleven nucleotides from the hairpin base (two helical RNA turns into the stem). The resulting hairpin, known as a pre-miRNA, has a two-nucleotide overhang at its 3’ end; it has 3' hydroxyl and 5' phosphate groups.
pre-miRNAs that are spliced directly out of introns, bypassing the Microprocessor complex, are known as "mirtrons." Originally thought to exist only in Drosophila and C. elegans, mirtrons have now been found in mammals.[46]
Perhaps as many as 16% of pri-miRNAs may be altered through nuclear RNA editing.[47][48][49] Most commonly, enzymes known as adenosine deaminases acting on RNA (ADARs) catalyze adenosine to inosine (A to I) transitions. RNA editing can halt nuclear processing (for example, of pri-miR-142, leading to degradation by the ribonuclease Tudor-SN) and alter downstream processes including cytoplasmic miRNA processing and target specificity (e.g., by changing the seed region of miR-376 in the central nervous system).[47]
Nuclear export
pre-miRNA hairpins are exported from the nucleus in a process involving the nucleocytoplasmic shuttle Exportin-5. This protein, a member of the karyopherin family, recognizes a two-nucleotide overhang left by the RNase III enzyme Drosha at the 3' end of the pre-miRNA hairpin. Exportin-5-mediated transport to the cytoplasm is energy-dependent, using GTP bound to the Ran protein.[50]
Cytoplasmic processing
In the cytoplasm, the pre-miRNA hairpin is cleaved by the RNase III enzyme Dicer.[51] This endoribonuclease interacts with the 3' end of the hairpin and cuts away the loop joining the 3' and 5' arms, yielding an imperfect miRNA:miRNA* duplex about 22 nucleotides in length.[51] Overall hairpin length and loop size influence the efficiency of Dicer processing, and the imperfect nature of the miRNA:miRNA* pairing also affects cleavage.[51][52] Although either strand of the duplex may potentially act as a functional miRNA, only one strand is usually incorporated into the RNA-induced silencing complex (RISC) where the miRNA and its mRNA target interact.
The RNA-induced silencing complex
Main article: RNA-induced silencing complex
The mature miRNA is part of an active RNA-induced silencing complex (RISC) containing Dicer and many associated proteins.[53] RISC is also known as a microRNA ribonucleoprotein complex (miRNP);[54] RISC with incorporated miRNA is sometimes referred to as "miRISC."
Dicer processing of the pre-miRNA is thought to be coupled with unwinding of the duplex. Generally, only one strand is incorporated into the miRISC, selected on the basis of its thermodynamic instability and weaker base-pairing relative to the other strand.(Krol et al., 2004)(Khvorova et al., 2003; Schwarz et al., 2003) The position of the stem-loop may also influence strand choice. (Lin et al., 2005) The other strand, called the passenger strand due to its lower levels in the steady state, is denoted with an asterisk (*) and is normally degraded. In some cases, both strands of the duplex are viable and become functional miRNA that target different mRNA populations.(Okamura et al., 2008)
Members of the argonaute (Ago) protein family are central to RISC function. Argonautes are needed for miRNA-induced silencing and contain two conserved RNA binding domains: a PAZ domain that can bind the single stranded 3’ end of the mature miRNA and a PIWI domain that structurally resembles ribonuclease-H and functions to interact with the 5’ end of the guide strand. They bind the mature miRNA and orient it for interaction with a target mRNA. Some argonautes, for example human Ago2, cleave target transcripts directly; argonautes may also recruit additional proteins to achieve translational repression.[55] The human genome encodes eight argonaute proteins divided by sequence similarities into two families: AGO (with four members present in all mammalian cells and called E1F2C/hAgo in humans), and PIWI (found in the germ line and hematopoietic stem cells).[55][56]
Additional RISC components include TRBP [human immunodeficiency virus (HIV) transactivating response RNA (TAR) binding protein],[57] PACT (protein activator of the interferon induced protein kinase (PACT), the SMN complex, fragile X mental retardation protein (FMRP), and Tudor staphylococcal nuclease-domain-containing protein (Tudor-SN).[58][59]
miRNA turnover
Turnover of mature miRNA is needed for rapid changes in miRNA expression profiles. During miRNA maturation in the cytoplasm, uptake by the Argonaute protein is thought to stabilize the guide strand, while the opposite (* or "passenger") strand is preferentially destroyed. In what has been called a "Use it or lose it" strategy, Argonaute may preferentially retain miRNAs with many targets over miRNAs with few or no targets, leading to degradation of the non-targeting molecules.[60]
Decay of mature miRNAs in animals is mediated by the 5´-to-3´ exoribonuclease XRN2, also known as Rat1p.[61] In plants, SDN (small RNA degrading nuclease) family members degrade miRNAs in the opposite (3'-to-5') direction. Similar enzymes are encoded in animal genomes, but their roles have not yet been described.[60]
Several miRNA modifications affect miRNA stability. As indicated by work in the model organism Arabidopsis thaliana (thale cress), mature plant miRNAs appear to be stabilized by the addition of methyl moieties at the 3' end. The 2'-O-conjugated methyl groups block the addition of uracil (U) residues by uridyltransferase enzymes, a modification that may be associated with miRNA degradation. However, uridylation may also protect some miRNAs; the consequences of this modification are incompletely understood. Uridylation of some animal miRNAs has also been reported. Both plant and animal miRNAs may be altered by addition of adenine (A) residues to the 3' end of the miRNA. An extra A added to the end of mammalian miR-122, a liver-enriched miRNA important in Hepatitis C, stabilizes the molecule, and plant miRNAs ending with an adenine residue have slower decay rates.[60]
Cellular functions
The function of miRNAs appears to be in gene regulation. For that purpose, a miRNA is complementary to a part of one or more messenger RNAs (mRNAs). Animal miRNAs are usually complementary to a site in the 3' UTR whereas plant miRNAs are usually complementary to coding regions of mRNAs.[62] Perfect or near perfect base pairing with the target RNA promotes cleavage of the RNA.[63] This is the primary mode of plant microRNAs.[64] In animals, microRNAs more often only partially base pair and inhibit protein translation of the target mRNA[65] (this exists in plants as well but is less common).[64] MicroRNAs that are partially complementary to the target can also speed up deadenylation, causing mRNAs to be degraded sooner.[66] For partially complementary microRNA to recognise their targets, the nucleotides 2–7 of the miRNA ('seed region') still have to be perfectly complementary.[67] miRNAs occasionally also causes histone modification and DNA methylation of promoter sites and therefore affecting the expression of targeted genes.[68][69]
Animal microRNAs target in particular developmental genes. In contrast, genes involved in functions common to all cells, such as gene expression, have very few microRNA target sites and seem to be under selection to avoid targeting by microRNAs.[70]
dsRNA can also activate gene expression, a mechanism that has been termed "small RNA-induced gene activation" or RNAa. dsRNAs targeting gene promoters can induce potent transcriptional activation of associated genes. This was demonstrated in human cells using synthetic dsRNAs termed small activating RNAs (saRNAs),[71] but has also been demonstrated for endogenous microRNA.[72]
Evolution
MicroRNAs are significant phylogenetic markers because of their astonishingly low rate of evolution.[73] Their origin may have permitted the development of morphological innovation, and by making gene expression more specific and 'fine-tunable', permitted the genesis of complex organs[74] and perhaps, ultimately, complex life.[75] Indeed, rapid bursts of morphological innovation are generally associated with a high rate of microRNA accumulation.[76][73]
MicroRNAs originate predominantly by the random formation of hairpins in "non-coding" sections of DNA (i.e. introns or intergene regions), but also by the duplication and modification of existing microRNAs.[77] The rate of evolution (i.e. nucleotide substitution) in recently-originated microRNAs is comparable to that elsewhere in the non-coding DNA, implying evolution by neutral drift; however, older microRNAs have a much lower rate of change (often less than one substitution per hundred million years),[75] suggesting that once a microRNA gains a function it undergoes extreme purifying selection.[77] At this point, a microRNA is rarely lost from an animal's genome,[75] although microRNAs which are more recently derived (and thus presumably non-functional) are frequently lost.[77] This makes them a valuable phylogenetic marker, and they are being looked upon as a possible solution to such outstanding phylogenetic problems as the relationships of arthropods.[78]
MicroRNAs feature in the genomes of most eukaryotic organisms, from the brown algae[79] to the metazoa. Across all species, in excess of 5000 had been identified by March 2010.[80] Whilst short RNA sequences (50 – hundreds of base pairs) of a broadly comparable function occur in bacteria, bacteria lack true microRNAs.[81]
Experimental detection and manipulation of miRNA
MicroRNA expression can be quantified in a two-step polymerase chain reaction process of modified RT-PCR followed by quantitative real-time PCR. Variations of this method achieve absolute or relative quantification.[82] miRNAs can also be hybridized to microarrays, slides or chips with probes to hundreds or thousands of miRNA targets, so that relative levels of miRNAs can be determined in different samples.[83] The activity of an miRNA can be experimentally inhibited using a locked nucleic acid oligo, a Morpholino oligo[84][85] or a 2'-O-methyl RNA oligo.[86] MicroRNA maturation can be inhibited at several points by steric-blocking oligos.[87] The miRNA target site of an mRNA transcript can also be blocked by a steric-blocking oligo.[88][89] Additionally, a specific miRNA can be silenced by a complementary antagomir.
miRNA and disease
Just as miRNA is involved in the normal functioning of eukaryotic cells, so has dysregulation of miRNA been associated with disease. A manually curated, publicly available database miR2Disease documents known relationships between miRNA dysregulation and human disease.[90]
miRNA and cancer
Several miRNAs have been found to have links with some types of cancer.[91][92]
A study of mice altered to produce excess c-Myc — a protein with mutated forms implicated in several cancers — shows that miRNA has an effect on the development of cancer. Mice that were engineered to produce a surplus of types of miRNA found in lymphoma cells developed the disease within 50 days and died two weeks later. In contrast, mice without the surplus miRNA lived over 100 days.[93] Leukemia can be caused by the insertion of a virus next to the 17-92 array of microRNAs leading to increased expression of this microRNA.[94]
Another study found that two types of miRNA inhibit the E2F1 protein, which regulates cell proliferation. miRNA appears to bind to messenger RNA before it can be translated to proteins that switch genes on and off.[95]
By measuring activity among 217 genes encoding miRNA, patterns of gene activity that can distinguish types of cancers can be discerned. miRNA signatures may enable classification of cancer. This will allow doctors to determine the original tissue type which spawned a cancer and to be able to target a treatment course based on the original tissue type. miRNA profiling has already been able to determine whether patients with chronic lymphocytic leukemia had slow growing or aggressive forms of the cancer.[96]
In the last few years, the experimental field that provided the deepest insights into the in vivo biology of miRNAs in cancer is that of mouse modeling, in which transgenic and knockout animals mimic over-expression or down-regulation, respectively, of specific miRNAs involved in human leukemia/lymphoma (Zanesi N et al, 2010).
Zanesi N, Pekarsky Y, Trapasso F, Calin G, Croce CM. MicroRNAs in mouse models of lymphoid malignancies. J Nucl Acid Invest, 1:e8, 36-40, 2010.
miRNA and heart disease
The global role of miRNA function in the heart has been addressed by conditionally inhibiting miRNA maturation in the murine heart, and has revealed that miRNAs play an essential role during its development.[97][98] miRNA expression profiling studies demonstrate that expression levels of specific miRNAs change in diseased human hearts, pointing to their involvement in cardiomyopathies.[99][100][101] Furthermore, studies on specific miRNAs in animal models have identified distinct roles for miRNAs both during heart development and under pathological conditions, including the regulation of key factors important for cardiogenesis, the hypertrophic growth response, and cardiac conductance.[98][102][103][104][105][106]
miRNA and the nervous system
miRNAs appear to regulate the nervous system.[107] Neural miRNAs are involved at various stages of synaptic development, including dendritogenesis, synapse formation and synapse maturation.[108] Some studies find altered miRNA expression in schizophrenia.[109][110]
miRNA and non-coding RNAs
When the human genome project mapped its first chromosome in 1999, it was predicted it would contain over 100,000 protein coding genes. However, only around 20,000 were eventually identified (International Human Genome Sequencing Consortium, 2004) and for a long time much of the non-protein-coding DNA was considered "junk", though conventional wisdom holds that much if not most of the genome is functional.[111] Since then, the advent of sophisticated bioinformatics approaches combined with genome tiling studies examining the transcriptome,[112] systematic sequencing of full length cDNA libraries,[113] and experimental validation[114] (including the creation of miRNA derived antisense oligonucleotides called antagomirs) have revealed that many transcripts are for non protein-coding RNA, including snoRNA and miRNA.[115]
See also
* Gene expression
* RNAi
* siRNA
* List of miRNA target prediction tools
* List of miRNA gene prediction tools
References
1. ^ Bartel, DP (2009). "MicroRNAs: target recognition and regulatory functions". Cell 136 (2): 215–33. doi:10.1016/j.cell.2009.01.002. PMID 19167326. edit
2. ^ Bartel, D. P. (2004). MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281-297.
3. ^ Homo sapiens miRNAs in the miRBase at Manchester University
4. ^ Bentwich, I., Avniel, A., Karov, Y., Aharonov, R., Gilad, S., Barad, O., Barzilai, A., Einat, P., Einav, U., Meiri, E., et al. (2005). Identification of hundreds of conserved and nonconserved human microRNAs. Nat Genet 37, 766-770.
5. ^ Friedman, R. C., Farh, K. K., Burge, C. B., and Bartel, D. P. (2009). Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 19, 92-105.
6. ^ Lim, L. P., Lau, N. C., Weinstein, E. G., Abdelhakim, A., Yekta, S., Rhoades, M. W., Burge, C. B., and Bartel, D. P. (2003). The microRNAs of Caenorhabditis elegans. Genes Dev 17, 991-1008.
7. ^ Brennecke, J., Stark, A., Russell, R. B., and Cohen, S. M. (2005). Principles of microRNA-target recognition. PLoS Biol 3, e85.
8. ^ Lim, L. P., Lau, N. C., Garrett-Engele, P., Grimson, A., Schelter, J. M., Castle, J., Bartel, D. P., Linsley, P. S., and Johnson, J. M. (2005). Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433, 769-773.
9. ^ Tanzer, A., and Stadler, P. F. (2004). Molecular evolution of a microRNA cluster. J Mol Biol 339, 327-335.
10. ^ Molnar, A., Schwach, F., Studholme, D. J., Thuenemann, E. C., and Baulcombe, D. C. (2007). miRNAs control gene expression in the single-cell alga Chlamydomonas reinhardtii. Nature 447, 1126-1129.
11. ^ Kren, B. T., Wong, P. Y., Sarver, A., Zhang, X., Zeng, Y., and Steer, C. J. (2009). microRNAs identified in highly purified liver-derived mitochondria may play a role in apoptosis. RNA Biol 6.
12. ^ Grosshans, H., and Slack, F. J. (2002). Micro-RNAs: small is plentiful. J Cell Biol 156, 17-21.
13. ^ Lee, C. T., Risom, T., and Strauss, W. M. (2007a). Evolutionary conservation of microRNA regulatory circuits: an examination of microRNA gene complexity and conserved microRNA-target interactions through metazoan phylogeny. DNA Cell Biol 26, 209-218.
14. ^ Brennecke, J., Hipfner, D. R., Stark, A., Russell, R. B., and Cohen, S. M. (2003). bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in Drosophila. Cell 113, 25-36.
15. ^ Cuellar, T. L., and McManus, M. T. (2005). MicroRNAs and endocrine biology. J Endocrinol 187, 327-332.
16. ^ Poy, M. N., Eliasson, L., Krutzfeldt, J., Kuwajima, S., Ma, X., Macdonald, P. E., Pfeffer, S., Tuschl, T., Rajewsky, N., Rorsman, P., and Stoffel, M. (2004). A pancreatic islet-specific microRNA regulates insulin secretion. Nature 432, 226-230.
17. ^ Chen, C. Z., Li, L., Lodish, H. F., and Bartel, D. P. (2004). MicroRNAs modulate hematopoietic lineage differentiation. Science 303, 83-86.
18. ^ Wilfred, B. R., Wang, W. X., and Nelson, P. T. (2007). Energizing miRNA research: a review of the role of miRNAs in lipid metabolism, with a prediction that miR-103/107 regulates human metabolic pathways. Mol Genet Metab 91, 209-217.
19. ^ Harfe, B. D., McManus, M. T., Mansfield, J. H., Hornstein, E., and Tabin, C. J. (2005). The RNaseIII enzyme Dicer is required for morphogenesis but not patterning of the vertebrate limb. Proc Natl Acad Sci U S A 102, 10898-10903.
20. ^ Lim, L. P., Lau, N. C., Garrett-Engele, P., Grimson, A., Schelter, J. M., Castle, J., Bartel, D. P., Linsley, P. S., and Johnson, J. M. (2005). Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433, 769-773.
21. ^ Lagos-Quintana, M., Rauhut, R., Meyer, J., Borkhardt, A., and Tuschl, T. (2003). New microRNAs from mouse and human. Rna 9, 175-179.
22. ^ Lagos-Quintana, M., Rauhut, R., Yalcin, A., Meyer, J., Lendeckel, W., and Tuschl, T. (2002). Identification of tissue-specific microRNAs from mouse. Curr Biol 12, 735-739.
23. ^ Trang, P; Weidhaas, JB; Slack, FJ (2008). "MicroRNAs as potential cancer therapeutics". Oncogene 27 Suppl 2: S52–7. doi:10.1038/onc.2009.353. PMID 19956180. edit
24. ^ Li, C; Feng, Y; Coukos, G; Zhang, L (2009). "Therapeutic microRNA strategies in human cancer". The AAPS journal 11 (4): 747–57. doi:10.1208/s12248-009-9145-9. PMID 19876744. edit
25. ^ Fasanaro, P.; Greco, S.; Ivan, M.; Capogrossi, M.; Martelli, F. (2010). "MicroRNA: emerging therapeutic targets in acute ischemic diseases". Pharmacology & therapeutics 125 (1): 92–104. doi:10.1016/j.pharmthera.2009.10.003. PMID 19896977. edit
26. ^ Lee, R. C., Feinbaum, R. L., and Ambros, V. (1993). The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75, 843-854.
27. ^ Reinhart, B. J., Slack, F. J., Basson, M., Pasquinelli, A. E., Bettinger, J. C., Rougvie, A. E., Horvitz, H. R., and Ruvkun, G. (2000). The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403, 901-906.
28. ^ Pasquinelli, A. E., Reinhart, B. J., Slack, F., Martindale, M. Q., Kuroda, M. I., Maller, B., Hayward, D. C., Ball, E. E., Degnan, B., Muller, P., et al. (2000). Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature 408, 86-89.
29. ^ Ambros, V., Bartel, B., Bartel, D. P., Burge, C. B., Carrington, J. C., Chen, X., Dreyfuss, G., Eddy, S. R., Griffiths-Jones, S., Marshall, M., et al. (2003). A uniform system for microRNA annotation. Rna 9, 277-279.
30. ^ Griffiths-Jones, S., Grocock, R. J., van Dongen, S., Bateman, A., and Enright, A. J. (2006). miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 34, D140-144.
31. ^ a b Lau, N. C., Lim, L. P., Weinstein, E. G., and Bartel, D. P. (2001). An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 294, 858-862.
32. ^ a b c d e f Lee, Y., Kim, M., Han, J., Yeom, K. H., Lee, S., Baek, S. H., and Kim, V. N. (2004). MicroRNA genes are transcribed by RNA polymerase II. Embo J 23, 4051-4060.
33. ^ Lagos-Quintana, M., Rauhut, R., Lendeckel, W., and Tuschl, T. (2001). Identification of novel genes coding for small expressed RNAs. Science 294, 853-858.
34. ^ Lee, R. C., and Ambros, V. (2001). An extensive class of small RNAs in Caenorhabditis elegans. Science 294, 862-864.
35. ^ Rodriguez, A., Griffiths-Jones, S., Ashurst, J. L., and Bradley, A. (2004). Identification of mammalian microRNA host genes and transcription units. Genome Res 14, 1902-1910.
36. ^ a b c d Cai, X., Hagedorn, C. H., and Cullen, B. R. (2004). Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. Rna 10, 1957-1966.
37. ^ Weber, M. J. (2005). New human and mouse microRNA genes found by homology search. Febs J 272, 59-73.
38. ^ Kim, Y. K., and Kim, V. N. (2007). Processing of intronic microRNAs. Embo J 26, 775-783.
39. ^ Baskerville, S., and Bartel, D. P. (2005). Microarray profiling of microRNAs reveals frequent coexpression with neighboring miRNAs and host genes. Rna 11, 241-247.
40. ^ Rodriguez, A., Griffiths-Jones, S., Ashurst, J. L., and Bradley, A. (2004). Identification of mammalian microRNA host genes and transcription units. Genome Res 14, 1902-1910.
41. ^ Altuvia, Y., Landgraf, P., Lithwick, G., Elefant, N., Pfeffer, S., Aravin, A., Brownstein, M. J., Tuschl, T., and Margalit, H. (2005). Clustering and conservation patterns of human microRNAs. Nucleic Acids Res 33, 2697-2706.
42. ^ Zhou, X., Ruan, J., Wang, G., and Zhang, W. (2007). Characterization and identification of microRNA core promoters in four model species. PLoS Comput Biol 3, e37.
43. ^ Zhou, X., Ruan, J., Wang, G., and Zhang, W. (2007). Characterization and identification of microRNA core promoters in four model species. PLoS Comput Biol 3, e37.
44. ^ Faller, M.; Guo, F. (2008). "MicroRNA biogenesis: there's more than one way to skin a cat.". Biochimica et biophysica acta 1779 (11): 663–667. doi:10.1016/j.bbagrm.2008.08.005. PMID 18778799. edit
45. ^ Gregory, R.; Chendrimada, T.; Shiekhattar, R. (2006). "MicroRNA biogenesis: isolation and characterization of the microprocessor complex". Methods in molecular biology (Clifton, N.J.) 342: 33–47. doi:10.1385/1-59745-123-1:33. PMID 16957365. edit
46. ^ Berezikov, E.; Chung, W.; Willis, J.; Cuppen, E.; Lai, E. (2007). "Mammalian mirtron genes". Molecular cell 28 (2): 328–336. doi:10.1016/j.molcel.2007.09.028. PMID 17964270. edit
47. ^ a b Kawahara, Y.; Megraw, M.; Kreider, E.; Iizasa, H.; Valente, L.; Hatzigeorgiou, A.; Nishikura, K. (2008). "Frequency and fate of microRNA editing in human brain". Nucleic acids research 36 (16): 5270–5280. doi:10.1093/nar/gkn479. PMID 18684997. edit
48. ^ Winter, J.; Jung, S.; Keller, S.; Gregory, R.; Diederichs, S. (2009). "Many roads to maturity: microRNA biogenesis pathways and their regulation". Nature cell biology 11 (3): 228–234. doi:10.1038/ncb0309-228. PMID 19255566. edit
49. ^ Ohman, M. (2007). "A-to-I editing challenger or ally to the microRNA process". Biochimie 89 (10): 1171–1176. doi:10.1016/j.biochi.2007.06.002. PMID 17628290. edit
50. ^ Murchison, E.; Hannon, G. (2004). "MiRNAs on the move: miRNA biogenesis and the RNAi machinery". Current opinion in cell biology 16 (3): 223–229. doi:10.1016/j.ceb.2004.04.003. PMID 15145345. edit
51. ^ a b c Lund, E.; Dahlberg, J. (2006). "Substrate selectivity of exportin 5 and Dicer in the biogenesis of microRNAs". Cold Spring Harbor symposia on quantitative biology 71: 59–66. doi:10.1101/sqb.2006.71.050. PMID 17381281. edit
52. ^ Ji, X (2008). "The mechanism of RNase III action: how dicer dices". Current topics in microbiology and immunology 320: 99–116. doi:10.1007/978-3-540-75157-1_5. PMID 18268841. edit
53. ^ Rana, T. M. (2007). "Illuminating the silence: understanding the structure and function of small RNAs". Nature Reviews Molecular Cell Biology 8 (1): 23. doi:10.1038/nrm2085. PMID 17183358. edit
54. ^ Schwarz, D.; Zamore, P. (2002). "Why do miRNAs live in the miRNP?". Genes & development 16 (9): 1025–1031. doi:10.1101/gad.992502. PMID 12000786. edit
55. ^ a b Pratt, A.; MacRae, I. (2009). "The RNA-induced silencing complex: a versatile gene-silencing machine". The Journal of biological chemistry 284 (27): 17897–17901. doi:10.1074/jbc.R900012200. PMID 19342379. edit
56. ^ Schwarz, D.; Zamore, P. (2002). "Why do miRNAs live in the miRNP?". Genes & development 16 (9): 1025–1031. doi:10.1101/gad.992502. PMID 12000786. edit
57. ^ MacRae, I.; Ma, E.; Zhou, M.; Robinson, C.; Doudna, J. (2008). "In vitro reconstitution of the human RISC-loading complex". Proceedings of the National Academy of Sciences of the United States of America 105 (2): 512–517. doi:10.1073/pnas.0710869105. PMID 18178619. edit
58. ^ Murchison, E.; Hannon, G. (2004). "MiRNAs on the move: miRNA biogenesis and the RNAi machinery". Current opinion in cell biology 16 (3): 223–229. doi:10.1016/j.ceb.2004.04.003. PMID 15145345. edit
59. ^ Mourelatos Z, Dostie J, Paushkin S, et al. (March 2002). "miRNPs: a novel class of ribonucleoproteins containing numerous microRNAs". Genes Dev. 16 (6): 720–8. doi:10.1101/gad.974702. PMID 11914277.
60. ^ a b c Kai, Z.; Pasquinelli, A. (2010). "MicroRNA assassins: factors that regulate the disappearance of miRNAs". Nature structural & molecular biology 17 (1): 5–10. doi:10.1038/nsmb.1762. PMID 20051982. edit
61. ^ Chatterjee S, Großhans H (September 2009). "Active turnover modulates mature microRNA activity in Caenorhabditis elegans". Nature 461 (7263): 546–459. doi:10.1038/nature08349. PMID 19734881.
62. ^ Wang XJ, Reyes JL, Chua NH, Gaasterland T (2004). "Prediction and identification of Arabidopsis thaliana microRNAs and their mRNA targets". Genome Biol. 5 (9): R65. doi:10.1186/gb-2004-5-9-r65. PMID 15345049. PMC 522872. http://genomebiology.com/2004/5/9/R65.
63. ^ Kawasaki H, Taira K (2004). "MicroRNA-196 inhibits HOXB8 expression in myeloid differentiation of HL60 cells". Nucleic Acids Symp Ser 48 (48): 211–2. doi:10.1093/nass/48.1.211. PMID 17150553.
64. ^ a b Moxon S, Jing R, Szittya G, et al. (October 2008). "Deep sequencing of tomato short RNAs identifies microRNAs targeting genes involved in fruit ripening". Genome Res. 18 (10): 1602–9. doi:10.1101/gr.080127.108. PMID 18653800.
65. ^ Williams AE (February 2008). "Functional aspects of animal microRNAs". Cell. Mol. Life Sci. 65 (4): 545–62. doi:10.1007/s00018-007-7355-9. PMID 17965831.
66. ^ Eulalio A, Huntzinger E, Nishihara T, Rehwinkel J, Fauser M, Izaurralde E (January 2009). "Deadenylation is a widespread effect of miRNA regulation". RNA 15 (1): 21–32. doi:10.1261/rna.1399509. PMID 19029310.
67. ^ Mazière P, Enright AJ (June 2007). "Prediction of microRNA targets". Drug Discov. Today 12 (11-12): 452–8. doi:10.1016/j.drudis.2007.04.002. PMID 17532529.
68. ^ Tan Y, Zhang B, Wu T, et al. (February 2009). "Transcriptional inhibition of Hoxd4 expression by noncoding RNAs in human breast cancer cells". BMC Mol. Biol. 10 (1): 12. doi:10.1186/1471-2199-10-12. PMID 19232136.
69. ^ Hawkins PG, Morris KV (March 2008). "RNA and transcriptional modulation of gene expression". Cell Cycle 7 (5): 602–7. PMID 18256543. PMC 2877389. http://www.landesbioscience.com/journals/cc/abstract.php?id=5522.
70. ^ Stark A, Brennecke J, Bushati N, Russell RB, Cohen SM (2005). "Animal MicroRNAs confer robustness to gene expression and have a significant impact on 3'UTR evolution". Cell 123 (6): 1133–46. doi:10.1016/j.cell.2005.11.023. PMID 16337999.
71. ^ Li LC (2008). "Small RNA-Mediated Gene Activation". RNA and the Regulation of Gene Expression: A Hidden Layer of Complexity. Caister Academic Press. ISBN 978-1-904455-25-7. http://www.horizonpress.com/rnareg.
72. ^ Place RF, Li LC, Pookot D, Noonan EJ, Dahiya R (2008). "MicroRNA-373 induces expression of genes with complementary promoter sequences". Proc. Natl. Acad. Sci. U.S.A. 105 (5): 1608–13. doi:10.1073/pnas.0707594105. PMID 18227514.
73. ^ a b Wheeler, B.; Heimberg, A.; Moy, V.; Sperling, E.; Holstein, T.; Heber, S.; Peterson, K. (2009). "The deep evolution of metazoan microRNAs". Evolution & development 11 (1): 50–68. doi:10.1111/j.1525-142X.2008.00302.x. PMID 19196333. edit
74. ^ Heimberg, A.; Sempere, L.; Moy, V.; Donoghue, P.; Peterson, K. (2008). "MicroRNAs and the advent of vertebrate morphological complexity". Proceedings of the National Academy of Sciences of the United States of America 105 (8): 2946–2950. doi:10.1073/pnas.0712259105. PMID 18287013. edit
75. ^ a b c Peterson, K.; Dietrich, M.; McPeek, M. (2009). "MicroRNAs and metazoan macroevolution: insights into canalization, complexity, and the Cambrian explosion". BioEssays : news and reviews in molecular, cellular and developmental biology 31 (7): 736–747. doi:10.1002/bies.200900033. PMID 19472371. edit
76. ^ Heimberg, A.; Sempere, L.; Moy, V.; Donoghue, P.; Peterson, K. (2008). "MicroRNAs and the advent of vertebrate morphological complexity". Proceedings of the National Academy of Sciences of the United States of America 105 (8): 2946–2950. doi:10.1073/pnas.0712259105. PMID 18287013. edit
77. ^ a b c Nozawa, M.; Miura, S.; Nei, M. (2010). "Origins and Evolution of MicroRNA Genes in Drosophila Species". Genome Biology and Evolution 2010: 180. doi:10.1093/gbe/evq009. edit
78. ^ Caravas, J.; Friedrich, M. (2010). "Of mites and millipedes: Recent progress in resolving the base of the arthropod tree". BioEssays 32 (6): 488. doi:10.1002/bies.201000005. PMID 20486135. edit
79. ^ Cock, J. M.; Sterck, L.; Rouzé, P.; Scornet, D.; Allen, A. E.; Amoutzias, G.; Anthouard, V.; Artiguenave, F. O. et al. (2010). "The Ectocarpus genome and the independent evolution of multicellularity in brown algae". Nature 465 (7298): 617–621. doi:10.1038/nature09016. PMID 20520714. edit
80. ^ Dimond, Patricia F. (15 March 2010), "miRNAs' Therapeutic Potential", Genetic Engineering & Biotechnology News 30 (6): 1, archived from the original on 10 July 2010, http://www.webcitation.org/5r7L7BfnG, retrieved 10 July 2010
81. ^ Tjaden, B.; Goodwin, S. S.; Opdyke, J. A.; Guillier, M.; Fu, D. X.; Gottesman, S.; Storz, G. (2006). "Target prediction for small, noncoding RNAs in bacteria". Nucleic Acids Research 34 (9): 2791. doi:10.1093/nar/gkl356. PMID 16717284. edit
82. ^ Caifu, Chen; Dana A. Ridzon, Adam J. Broomer, Zhaohui Zhou, Danny H. Lee, Julie T. Nguyen, Maura Barbisin, Nan Lan Xu, Vikram R. Mahuvakar, Mark R. Andersen, Kai Qin Lao, Kenneth J. Livak and Karl J. Guegler (2005-10-25). "Real-time quantification of microRNAs by stem–loop RT–PCR". Nucleic Acids Research 33 (20): e179. doi:10.1093/nar/gni178. PMID 16314309.
83. ^ Jaclyn, Shingara; KERRI KEIGER, JEFFREY SHELTON, WALAIRAT LAOSINCHAI-WOLF, PATRICIA POWERS, RICHARD CONRAD, DAVID BROWN, EMMANUEL LABOURIER (2005-07-25). "An optimized isolation and labeling platform for accurate microRNA expression profiling". RNA 11 (9): 1461–1470. doi:10.1261/rna.2610405. PMID 16043497.
84. ^ Kloosterman WP, Wienholds E, Ketting RF, Plasterk RH (2004). "Substrate requirements for let-7 function in the developing zebrafish embryo". Nucleic Acids Res. 32 (21): 6284–91. doi:10.1093/nar/gkh968. PMID 15585662.
85. ^ Flynt AS, Li N, Thatcher EJ, Solnica-Krezel L, Patton JG (February 2007). "Zebrafish miR-214 modulates Hedgehog signaling to specify muscle cell fate". Nat. Genet. 39 (2): 259–63. doi:10.1038/ng1953. PMID 17220889.
86. ^ Meister G, Landthaler M, Dorsett Y, Tuschl T (March 2004). "Sequence-specific inhibition of microRNA- and siRNA-induced RNA silencing". RNA 10 (3): 544–50. doi:10.1261/rna.5235104. PMID 14970398.
87. ^ Kloosterman WP, Lagendijk AK, Ketting RF, Moulton JD, Plasterk RH (August 2007). "Targeted inhibition of miRNA maturation with morpholinos reveals a role for miR-375 in pancreatic islet development". PLoS Biol. 5 (8): e203. doi:10.1371/journal.pbio.0050203. PMID 17676975.
88. ^ Choi, WY; Giraldez AJ, Schier AF (2007). "Target Protectors Reveal Dampening and Balancing of Nodal Agonist and Antagonist by miR-430." (Pubmed). Science. 318 (5848): 271–4. doi:10.1126/science.1147535. PMID 17761850.
89. ^ Klein, E.; Lioy, T.; Ma, L.; Impey, S.; Mandel, G.; Goodman, H. (Dec 2007). "Homeostatic regulation of MeCP2 expression by a CREB-induced microRNA". Nature neuroscience 10 (12): 1513–1514. doi:10.1038/nn2010. ISSN 1097-6256. PMID 17994015. edit
90. ^ Jiang Q, Wang Y, Hao Y, Juan L, Teng M, Zhang X, Li M, Wang G, Liu Y. (January 2009). "miR2Disease: a manually curated database for microRNA deregulation in human disease.". Nucleic Acids Research 37 (Database issue) (Database issue): D98–104. doi:10.1093/nar/gkn714. PMID 18927107.
91. ^ He L, Thomson JM, Hemann MT, et al. (June 2005). "A microRNA polycistron as a potential human oncogene". Nature 435 (7043): 828–33. doi:10.1038/nature03552. PMID 15944707.
92. ^ Mraz M, Pospisilova S, Malinova K, et al. (March 2009). "MicroRNAs in chronic lymphocytic leukemia pathogenesis and disease subtypes". Leuk Lymphoma 50 (3): 506–9. doi:10.1080/10428190902763517. PMID 19347736.
93. ^ He L, Thomson JM, Hemann MT, et al. (June 2005). "A microRNA polycistron as a potential human oncogene". Nature 435 (7043): 828–33. doi:10.1038/nature03552. PMID 15944707.
94. ^ Cui JW, Li YJ, Sarkar A, Brown J, Tan YH, Premyslova M, Michaud C, Iscove N, Wang GJ, Ben-David Y. (June 2007). "Retroviral insertional activation of the Fli-3 locus in erythroleukemias encoding a cluster of microRNAs that convert Epo-induced differentiation to proliferation.". Blood 110 (7): 2631–40. doi:10.1182/blood-2006-10-053850. PMID 17586726.
95. ^ O'Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT (June 2005). "c-Myc-regulated microRNAs modulate E2F1 expression". Nature 435 (7043): 839–43. doi:10.1038/nature03677. PMID 15944709.
96. ^ Lu J, Getz G, Miska EA, et al. (June 2005). "MicroRNA expression profiles classify human cancers". Nature 435 (7043): 834–8. doi:10.1038/nature03702. PMID 15944708.
97. ^ Chen JF, Murchison EP, Tang R, et al. (February 2008). "Targeted deletion of Dicer in the heart leads to dilated cardiomyopathy and heart failure". Proc. Natl. Acad. Sci. U.S.A. 105 (6): 2111–6. doi:10.1073/pnas.0710228105. PMID 18256189.
98. ^ a b Zhao Y, Ransom JF, Li A, et al. (April 2007). "Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2". Cell 129 (2): 303–17. doi:10.1016/j.cell.2007.03.030. PMID 17397913.
99. ^ Thum T, Galuppo P, Wolf C, et al. (July 2007). "MicroRNAs in the human heart: a clue to fetal gene reprogramming in heart failure". Circulation 116 (3): 258–67. doi:10.1161/CIRCULATIONAHA.107.687947. PMID 17606841.
100. ^ van Rooij E, Sutherland LB, Liu N, et al. (November 2006). "A signature pattern of stress-responsive microRNAs that can evoke cardiac hypertrophy and heart failure". Proc. Natl. Acad. Sci. U.S.A. 103 (48): 18255–60. doi:10.1073/pnas.0608791103. PMID 17108080.
101. ^ Tatsuguchi M, Seok HY, Callis TE, et al. (June 2007). "Expression of microRNAs is dynamically regulated during cardiomyocyte hypertrophy". J. Mol. Cell. Cardiol. 42 (6): 1137–41. doi:10.1016/j.yjmcc.2007.04.004. PMID 17498736.
102. ^ Zhao Y, Samal E, Srivastava D (July 2005). "Serum response factor regulates a muscle-specific microRNA that targets Hand2 during cardiogenesis". Nature 436 (7048): 214–20. doi:10.1038/nature03817. PMID 15951802.
103. ^ Xiao J, Luo X, Lin H, et al. (April 2007). "MicroRNA miR-133 represses HERG K+ channel expression contributing to QT prolongation in diabetic hearts". J. Biol. Chem. 282 (17): 12363–7. doi:10.1074/jbc.C700015200. PMID 17344217. http://www.jbc.org/cgi/pmidlookup?view=long&pmid=17344217.
104. ^ Yang B, Lin H, Xiao J, et al. (April 2007). "The muscle-specific microRNA miR-1 regulates cardiac arrhythmogenic potential by targeting GJA1 and KCNJ2". Nat. Med. 13 (4): 486–91. doi:10.1038/nm1569. PMID 17401374.
105. ^ Carè A, Catalucci D, Felicetti F, et al. (May 2007). "MicroRNA-133 controls cardiac hypertrophy". Nat. Med. 13 (5): 613–8. doi:10.1038/nm1582. PMID 17468766.
106. ^ van Rooij E, Sutherland LB, Qi X, Richardson JA, Hill J, Olson EN (April 2007). "Control of stress-dependent cardiac growth and gene expression by a microRNA". Science (journal) 316 (5824): 575–9. doi:10.1126/science.1139089. PMID 17379774. http://www.sciencemag.org/cgi/pmidlookup?view=long&pmid=17379774.
107. ^ Maes OC, Chertkow HM, Wang E, Schipper HM. MicroRNA: implications for alzheimer disease and other human CNS disorders. Curr. Genomics 2009, 10 (3)154-168. PMID 19881909
108. ^ Schratt G. microRNAs at the synapse. Nat. Rev. Neurosc. 2009, 10 (12) 842-849. PMID 19888283
109. ^ Evidence for X-chromosomal schizophrenia associated with microRNA alterations. Feng J, Sun G, Yan J, Noltner K, Li W, Buzin CH, Longmate J, Heston LL, Rossi J, Sommer SS. PLoS One. 2009 Jul 1;4(7):e6121. PMID 19568434
110. ^ Beveridge NJ, Gardiner E, Carroll AP, Tooney, PA, Cairns MJ. Schizophrenia is associated with an increase in cortical microRNA biogenesis. Mol. Psychiatry 2009, Sep 1. PMID 19721432
111. ^ Pheasant, M., and Mattick, J. S. (2007). Raising the estimate of functional human sequences. Genome Res 17, 1245-1253.
112. ^ Bertone, P., Stolc, V., Royce, T. E., Rozowsky, J. S., Urban, A. E., Zhu, X., Rinn, J. L., Tongprasit, W., Samanta, M., Weissman, S., et al. (2004). Global identification of human transcribed sequences with genome tiling arrays. Science 306, 2242-2246.
113. ^ Ota, T., Suzuki, Y., Nishikawa, T., Otsuki, T., Sugiyama, T., Irie, R., Wakamatsu, A., Hayashi, K., Sato, H., Nagai, K., et al. (2004b). Complete sequencing and characterization of 21,243 full-length human cDNAs. Nat Genet 36, 40-45.
114. ^ Kuhn, D. E., Martin, M. M., Feldman, D. S., Terry, A. V., Jr., Nuovo, G. J., and Elton, T. S. (2008). Experimental validation of miRNA targets. Methods 44, 47-54.
115. ^ Huttenhofer, A., Schattner, P., and Polacek, N. (2005). Non-coding RNAs: hope or hype? Trends Genet 21, 289-297.
Further reading
* This paper discusses the role of microRNAs in viral oncogenesis: Scaria V; Jadhav, Vaibhav (2007). "microRNAs in viral oncogenesis.". Retrovirology 4 (82): 68. doi:10.1186/1742-4690-4-82. PMID 18036240.
* This paper discusses the role of microRNAs in Host-virus interactions: Scaria V; Hariharan, M; Maiti, S; Pillai, B; Brahmachari, SK (2006). "Host-Virus Interaction: A new role for microRNAs.". Retrovirology 3 (1): 68. doi:10.1186/1742-4690-3-68. PMID 17032463.
* This paper defines miRNA and proposes guidelines to follow in classifying RNA genes as miRNA: Ambros V, Bartel B, Bartel DP, Burge CB, Carrington JC, Chen X, Dreyfuss G, Eddy SR, Griffiths-Jones S, Marshall M, Matzke M, Ruvkun G, Tuschl T (2003). "A uniform system for microRNA annotation". RNA 9 (3): 277–279. doi:10.1261/rna.2183803. PMID 12592000.
* This paper discusses the processes that miRNA and siRNAs are involved in, in the context of 2 articles in the same issue of the journal Science: Baulcombe D (2002). "DNA events. An RNA microcosm.". Science 297 (5589): 2002–2003. doi:10.1126/science.1077906. PMID 12242426.
* This paper describes the discovery of lin-4, the first miRNA to be discovered: Lee RC, Feinbaum RL, Ambros V (1993). "The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14". Cell 75 (5): 843–854. doi:10.1016/0092-8674(93)90529-Y. PMID 8252621.
* "This paper showes that the over-expression of a miRNA in mice leads to cancer:" Costinean S, Zanesi N, Pekarsky Y, Tili E, Volinia S, Heerema N, Croce CM. Pre-B cell proliferation and lymphoblastic leukemia/high-grade lymphoma in E(mu)-miR155 transgenic mice. Proc Natl Acad Sci USA, 103(18): 7024-9, 2006.
External links
* The miRBase database
* The miRNA Blog
* miR2Disease, a manually curated database documenting known relationships between miRNA dysregulation and human disease.
Retrieved from "http://en.wikipedia.org/"
All text is available under the terms of the GNU Free Documentation License

